Keywords:
By Beatrice Marg-Haufe
Urban planning, urban warfare, urban decay..., and next up, urban metagenomics. If you had any doubt that we are living in the genomics era, consider this: On June 21st 2016, the International MetaSUB Consortium began collecting and mapping the genomes and epigenomes of microbial samples from 54 cities worldwide.
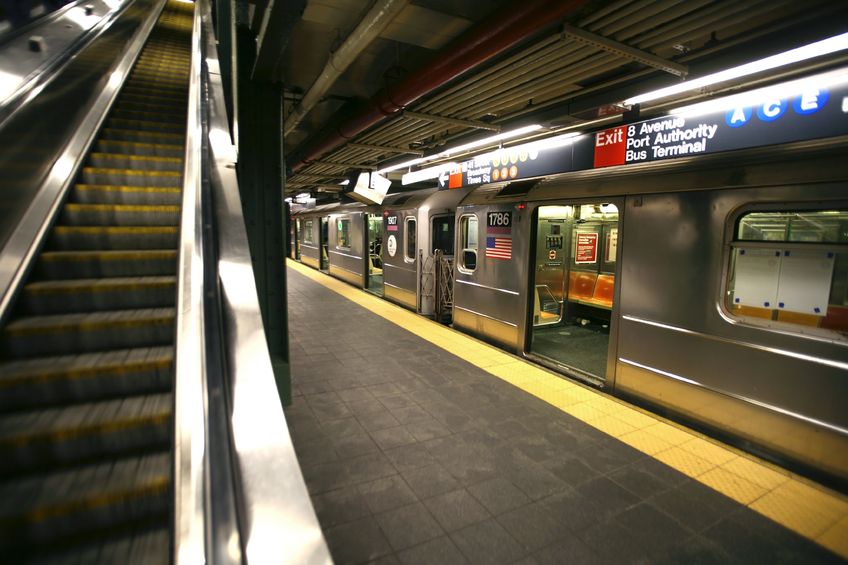

Behold the trillions of invisible inhabitants of New York City.
Supported by the Bill & Melinda Gates Foundation, the Alfred P. Sloan Foundation, Promega, Qiagen, OneCodes, and Illumina, MetaSUB (Metagenomics and Metadesign of Subways and Urban Biomes; www. metasub.org) aims to study the diversity of urban microbial communities and explore the genetic basis of antimicrobial resistance.
CosmosID, a genomic big data company that uses its curated genomic databases to identify microorganisms and detect antimicrobial resistance (AMR) genes in post-sequencing metagenomic samples, announced that it is working together with Weill Cornell Medicine to support the MetaSUB project and apply its technology to aid in microbial identification and the characterization of AMR genes.
Microbiome initiatives expand
The MetaSUB Consortium is testing various commercial kits for use in DNA and RNA extraction, including those of MoBio, Promega, Zymo, Omega BioTek, and Qiagen. All sample processing and shotgun sequencing using Illumina HiSeq technology will take place at regional MetaSUB hubs in New York City (for the Americas), Stockholm (Europe and Africa), and Shanghai (Asia and Oceania).
New metagenomics initiatives and research groups are forming all over the world. Companies, organizations, and governments are realizing the wide-ranging implications and potential applications of the discoveries being made about microbial populations – their diversity, function, and often precarious symbiotic existences. Technology advances that are improving and accelerating genomic workflows, including sample preparation and next-generation sequencing methodologies, are enabling large-scale metagenomic studies to be done more efficiently, requiring less time, labor, and costs.
The Microbiome Center is another recently formed partnership that combines the expertise and resources of The University of Chicago, the Marine Biological Laboratory (MBL), and the U.S. Department of Energy's Argonne National Laboratory. The Center will identify and seek to understand the function of bacteria, viruses, and fungi across environments. "The new center will broaden the look we have at the microbiome," said Huntington Willard, president and director of MBL. "By asking the same questions on different settings and scales, my guess is that we will discover similar principles at work from people studying inner city microbes and those in deep oceans - and that's where we can say something fundamental about how life works."
Resisting arrest: Antibiotic resistance genes
Antibiotics are physicians' weapons of choice to combat bacterial infections. However, clinicians' effective antimicrobial options are diminishing as bacteria continue to outsmart the pharmaceutical industry, developing new mechanisms of antibiotic resistance faster than companies can discover new classes of antimicrobial agents. Researchers are using metagenomics to profile and track the abundance and changes in antibiotic resistance genes within and across human populations. The information obtained from these studies will help scientists better understand the underlying mechanisms of resistance and develop new strategies, assays, and screens to identify novel compounds able to block antibiotic resistance.
In "Metagenome-wide Analysis of Antibiotic Resistance Gene in a Large Cohort of Human Gut Microbiota," for example, Hu et al. identified 1,093 antibiotic resistance genes in 162 individuals from China, Spain, and Denmark.1 The researchers showed that the antibiotic resistance genes from the two European populations were more closely related, whereas the genes from the Chinese samples tended to cluster separately. Overall, the samples from Chinese individuals had the highest number and abundance of antibiotic resistance genes, followed by those of Danish and Spanish individuals.
Forslund and coauthors studied the effects of antibiotic use on the human gut resistome by analyzing the metagenomic data obtained from stool samples using Illumina technology.2 To determine the impact of country-specific antibiotic consumption on the resistome, the researchers examined 252 fecal metagenomes from three countries and quantified the known resistance genes for 68 classes and subclasses of antibiotics. They found that the most abundant resistance genes correlate to those antibiotics used in food-producing animals and to antibiotics that have been available for a longer period of time. The authors concluded, "This outcome of our global, metagenomic-based approach, mapping variation within and between populations and covering a vast range of antibiotics, should provide a profound molecular basis for the ongoing debate on the appropriate use of antibiotics in agriculture and medicine."
Analyzing the metagenome
Next-generation sequencing is a powerful tool for metagenomic analysis. Automation of the sample processing, library preparation, and other steps in the NGS workflow can greatly increase throughput and reduce the cost of genomics research. Sample prep operations including DNA or RNA extraction and purification, preparation of library fragments, and PCR set-up can all introduce bottlenecks into the workflow and are all amenable to automation solutions.
The Freedom EVO® workstation from Tecan can automate most aspects of sample and library preparation for metagenomic analysis. Tecan works in collaboration with major research centers and NGS sample prep kit vendors – including Illumina® and Thermo Fisher Scientific – to develop and optimize automated NGS sample prep protocols.
FURTHER READING
- Hu Y, Yang X, Qin J, et al. Metagenome-wide analysis of antibiotic resistance genes in a large cohort of human gut microbiota. Nature Comm 2013;doi:10.1038/ncomms2151.
- Forslund K, Shinichi S, Kultima JR, et al. Country-specific Antibiotic Use Practices Impact the Human Gut Resistome. Genome Res 2013;23(7):1163-1169.
About the author

Dr Beatrice Marg-Haufe
Dr. Beatrice Marg-Haufe is a product manager at Tecan Switzerland with over 10 years of experience in assay development and product management. She studied biochemistry at the University of Bielefeld, Germany, and at Harvard Medical School, USA. She focused on cancer research during her PhD in Biochemistry at the MPI, Munich, Germany. She joined Tecan in 2009 focusing on applications for the agriculture and genomics market.