If your library preparation kit isn’t specifically designed for viral pathogen sequencing, then you’re probably missing data
Viral pathogens like SARS-CoV-2 can be detected by RT-PCR in nasal or throat swab samples from patients with COVID-19, but it is the detailed, sequence-based analysis that provides an understanding of the nature and mutation of the virus and the resulting disease state. Next generation sequencing (NGS) offers a comprehensive view of the complete genome, providing information about viral origins, pathogenesis, genetic variations, immune responses and phylogenetics.
Starting samples for pathogen sequencing are almost always of variable quality, with at least some samples degraded or low concentration, and some inevitably falling below the detection threshold of many current amplification technologies.
Highly sensitive detection and characterization of SARS-CoV-2 in nasal swab samples
with Revelo™ RNA-Seq library preparation kit
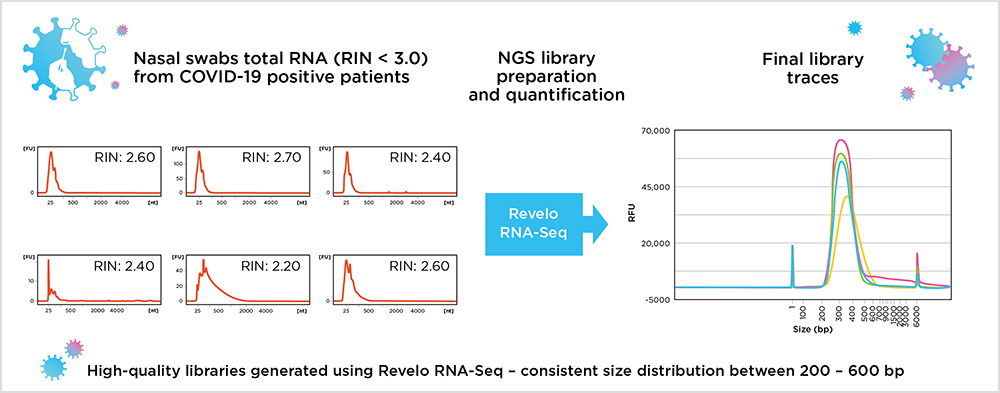
Performance of Revelo RNA-Seq in processing low-quality viral RNA from nasal swab samples
An upfront amplification method effectively addresses the challenge of ultra-low input to increase detection sensitivity, particularly in asymptomatic carriers. Our Single Primer Isothermal Amplification (SPIA®) technology has been used in hundreds of peer reviewed publications to enable access to low input RNA samples from various sources.
Key benefits
- Ability to obtain consistent results with low quality, degraded samples
- Effective DNase treatment to eliminate unwanted contaminating gDNA
- High detection sensitivity from samples with low viral titers in high host background
- Human rRNA depletion to enhance informative reads
Revelo RNA-Seq operates consistently across a very wide dynamic range and continues working at concentration levels far below the competitor’s detection threshold
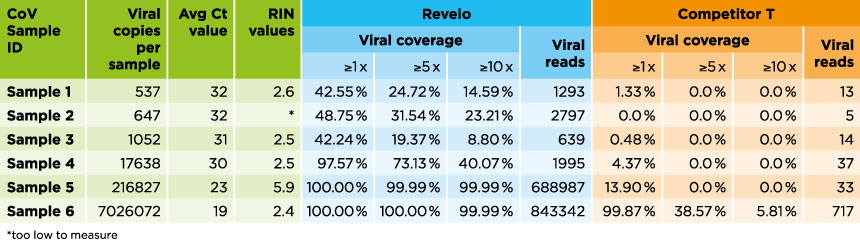
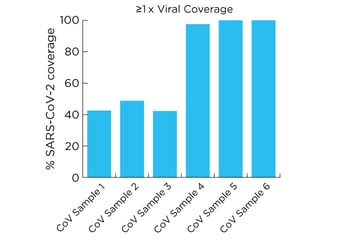
The detection sensitivity and coverage across a broad range of viral loads is shown here.
Results successfully demonstrate the high detection sensitivity of SARS-CoV-2 using Revelo RNA-Seq, from as low as ~500 viral copies.
For the competitor kit, viral coverage drops drastically, even when the viral copy number is as high as ~200,000.
Specifically designed for the amplification of pathogenic viral RNA from human samples, Revelo RNA-Seq library preparation kit enables a rapid six and half hours workflow from sample-to-sequencer.
For Research Use Only. Not for use in diagnostic procedures.