Keywords:
By Roberta Veneroni
Next-generation sequencing (NGS) is helping to advance genomics research at an unprecedented rate. However, the process can be technically challenging, and any errors can significantly impact the reliability and accuracy of your results. NGS library preparation and QC can have a major impact on your success, especially as poor-quality libraries can skew your results and reduce the accuracy of your data.
To help overcome these challenges, we’ve gathered our top five tips for optimizing your library preparation, quantification and QC processes, so you can produce better quality libraries, improve efficiency and reduce costs.
— Want to see how a novel quantification method can improve NGS library QC to save you time and money? Download our latest application note —
How can you overcome NGS library prep challenges to generate accurate data?
Tip 1: Choose appropriate NGS library prep reagents and kits
Choosing the right NGS library preparation kit can help you to streamline your process and generate high-quality libraries. Ideally, you should select kits that require the fewest steps, as this decreases the length of your library preparation process while minimizing sample loss and the likelihood of errors. One way to achieve this is to use kits that reduce the number of bead clean-up steps.
It’s also important to check whether the kit you choose is compatible with a wide range of input levels. Not all samples are equal, so it’s best to use kits that can deliver high-quality libraries regardless of the sample input range, rather than choosing a kit that is optimal for just one of your samples.
Another common issue that specialized kits can help overcome is the removal of rRNA when creating libraries for RNA sequencing. rRNA depletion is important as this type of RNA does not provide useful information and can dominate your sequencing data. As such, when performing RNA sequencing, we recommend selecting a kit that includes rRNA depletion after library preparation and can be customized to your species of interest.
Finally, as we explore in more detail below, automation can bring a number of benefits to your NGS workflows, but the approach is most effective when combined with kits designed to work with automated platforms. Select kits have been optimized on automated platforms to reduce the number of workflow steps and minimize manual input.
Tip 2: Automate your NGS workflows
Automated library preparation offers several advantages over manual NGS workflows, most notably when it comes to speed and efficiency. The latest automated systems can prepare samples quickly and on a large scale with minimal to no manual interaction. What’s more, it’s not just large laboratories that can benefit from automating routine, manual processes – any lab with limited resources and expanding workloads will benefit from freeing up trained scientists to work on more skilled tasks.
An additional benefit offered by automated NGS workflows is that they can improve the quality of your data. With manual workflows, differences in pipetting techniques between users can lead to significant variability in library yield. This can result in unnecessary costs and wasted resources as the libraries may have to be prepared again.
If you want to take advantage of these benefits, there’s a range of automated systems available for you to choose from. Traditionally, many of these tools only automate the part of the workflow leading up to library QC, but the most advanced can now automate all stages of library preparation and QC. The key advantage of integrating QC into your automated workflow is that you avoid manual normalization, which usually requires a number of calculations and complex pipetting steps. Integrated systems also make it easier and more convenient to troubleshoot issues when things go wrong, as you can lean on a single supplier to support your library, automation and QC needs.
Tip 3: QC all of your NGS libraries
If you want to optimize your NGS library preparation workflow, effective QC is crucial. So, which QC methods should you choose? There are a range of techniques in common use, including qPCR, microfluidic electrophoresis and fluorometry. However, microfluidic electrophoresis and fluorometry tend to offer low accuracy, as they only measure total nucleic acid concentration, meaning results can be influenced by non-sequenceable molecules. qPCR offers increased accuracy, but it is often more labor- and time-intensive, as it requires the preparation of several sample dilutions and replicates. This also introduces more variability and increases the chances of error.
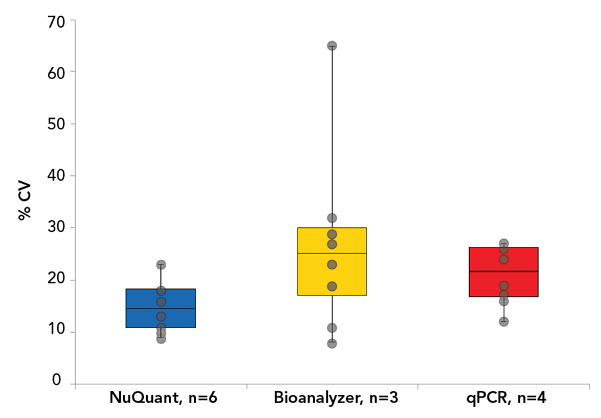
To solve some of these limitations, a novel quantification method known as NuQuant® has been developed. Using a simple fluorescent read-out, NuQuant offers high speed and accuracy, while eliminating the need for multiple manual steps (which also helps to reduce costs and minimize error). When compared to other methods, such as qPCR and microfluidic electrophoresis, NuQuant also delivers the lowest data variability from user to user, as shown in the figure below. You can learn more about NuQuant in this application note.
Tip 4: Minimize index hopping when preparing your NGS library
Index hopping occurs when one library’s barcode ends up on a different library such that the read is mis-assigned to the wrong sample, compromising the accuracy of your results. While this can have a major impact on any experiment, it’s especially important to minimize barcode hopping when high sensitivity is required on patterned flow cells.
Many new technologies now add unique dual index (UDI) pairs to libraries, ensuring that library management software will recognize barcode hopping when it occurs and remove any affected reads from your analysis. With these new solutions you can be more confident in the reliability of your data.
Tip 5: Prevent contamination during NGS library prep
Contamination is a big issue in NGS library preparation, especially when dealing with multiple samples. There are a number of things you can do to help avoid contamination. For manual methods, it’s crucial to adhere to good laboratory practice, such as sealing reaction tubes quickly, pipetting gently, handling samples carefully and wearing gloves. You should also always ensure your workspace is free of DNA and RNA.
When performing PCR, you should split your working space into at least two dedicated areas: one for pre-PCR preparation and one for post-PCR amplification steps and analysis. If your laboratory routinely conducts PCR, it might be worth considering segmenting these work areas further if you haven’t already.
However, even with the best precautions, manual methods will always be prone to contamination. Fully automated library preparation can solve this issue – as long as your sample is clean and your system is well-maintained, it’s far less likely that you will introduce contamination, giving you increased confidence in your results.
How to get started with improving your library prep workflow
Hopefully these expert tips will help you overcome any issues and challenges you are facing when it comes to NGS library prep. If you’re relatively new to optimizing your library preparation protocols and are wondering where you should start, we recommend beginning by reviewing and updating your QC processes – this can have a huge impact on the quality of your NGS data, while saving you time and money in the future.
For an in-depth exploration of how you can improve your NGS library QC processes, download our latest application note, which demonstrates a simple and accurate method for quantification and QC of your NGS libraries.
Keywords:
About the author

Roberta Veneroni
Dr. Roberta Veneroni is a Product and Applications Manager at Tecan with a focus in the field of Genomics. She has more than 10 years’ experience in developing assays and product management in the automation industry. She studied Biotechnology at the University of Milan, Italy, with orientation in cancer research in her PhD studies at the University of Eastern Piedmont in Novara, Italy. She has gained research experience at the UCI in the USA. When she joined the industry, she attended an international MBA at the ZHAW, Switzerland, focusing on globalization, multi-cultural environments and emerging markets. She joined Tecan in 2019 as responsible for the NGS liquid handling product portfolio.