By Enrique Neumann
Next-generation sequencing (NGS) has generated a raft of new developments and discoveries. However, NGS is a complex process, and scientists face many technical difficulties throughout the workflow. NGS sample preparation, for example, can be a significant source of inefficiencies that could hinder your research and stifle your progress by wasting resources and increasing costs. So, what can you do to improve your sample preparation efficiency? If you want to improve your NGS sample preparation process, a good place to start is NGS library prep. Library QC is usually a pain point, as even the most widely used QC methods suffer from a range of limitations in terms of time, cost and accuracy. Here we describe the options available to improve your library QC process, including a new method that could help you improve library quality while saving time and resources.

Improving your NGS library QC process is a powerful way to save time and increase data quality
Why you should optimize your NGS library QC process
Accurate library quantification and QC is critical for effective NGS, because the sequencing process relies on loading a very precise amount of sample onto the flow cell. If the concentration of your libraries is too high, the flow cell can become over-clustered and this can lead to run failure. On the other hand, if library concentration is too low, you’ll suffer from inefficiency and increased sequencing costs due to under-clustering. You can overcome these risks by employing effective QC processes to measure library concentration, because you can then normalize the concentration of all samples before they are pooled onto the sequencer.
Normalizing library concentration in this way is crucial to ensure efficiency when multiplexing libraries. This is because the number of reads per sample directly correlates with the sample molar concentration when libraries are pooled. As such, lower concentration libraries can be under-represented during this process, meaning that you may have to re-sequence your sample, which could waste valuable time and resources.
What’s more, these inefficiencies could have a growing impact on your research as your sequencing capacity expands – the more samples you run, the more work you may need to repeat. Since new high-capacity sequencers are now able to support the multiplexing of over 300 samples, it’s vital to look towards the future and cut down on your need for re-sequencing before scaling up your operations.
Given these issues, it’s essential to use effective QC processes to gain an accurate measure of library concentration that allows you to efficiently normalize your libraries and so achieve optimal yield and data quality. However, it can be a challenge to do this using the commonly used QC methods, as there is often a trade-off between accuracy and needing to save time and costs.
The limitations of common NGS library QC methods
Currently, the most frequently used NGS library QC techniques are qPCR, fluorometry and microfluidic electrophoresis. However, all of these methods have significant drawbacks in terms of accuracy, time or costs. For example, classic fluorometric methods only provide partial information, as they give concentration in ng/µl rather than molarity. They also suffer from low accuracy, as they only measure total nucleic acid concentration. Since this measurement will include non-sequenceable molecules (such as adapter dimers), quantification doesn’t give a reliable, representative measure of library concentration. In comparison, some electrophoresis methods are capable of better precision, but they are costly and slow, especially when checking samples individually.
qPCR is the current gold standard, as it only quantifies the molecules of interest and therefore has much better accuracy. However, the approach has other drawbacks. The qPCR process is labor- and time-intensive, involving several sample dilutions and requiring an initial stage of fragment size analysis beforehand. These multiple, manual steps can introduce error and user-user variability, leading to problems with the consistency and reliability of results. What’s more, qPCR is relatively expensive in terms of the reagent kits and consumables required. Given the multiple limitations of qPCR, there’s a real need for alternative QC processes that overcome these drawbacks in terms of time, cost and variability.
How you can overcome these limitations to optimize your NGS library QC process
A novel quantification method known as NuQuant® can now overcome these limitations. Integrated into select DNA-Seq and mRNA-Seq library preparation kits, NuQuant enables highly accurate and consistent NGS library QC at significantly reduced time and cost. The principle of the NuQuant method is to incorporate a specific number of fluorescent labels into each library molecule, regardless of its size, so that library molar concentration can be directly measured by fluorometry without the need for separate fragment size analysis. This capacity for direct measurement makes NuQuant the simplest QC method currently available.
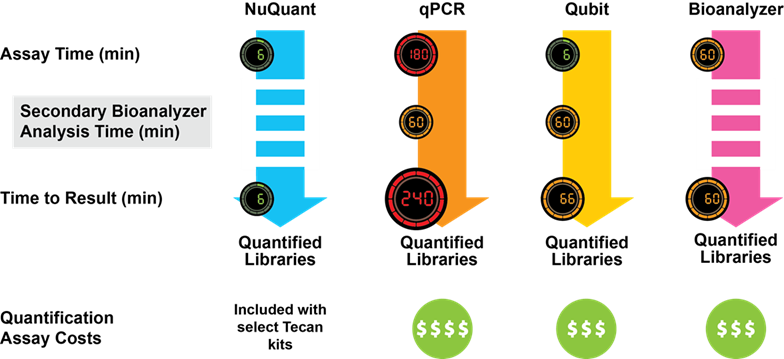
The streamlined design of the NuQuant method gives it several key advantages over other QC techniques. Firstly, it is much more time- and cost-efficient. In fact, the time savings you can make are quite astonishing – the entire workflow can be completed in under six minutes, whereas other methods take between one and four hours. Much of this difference comes from eliminating the need for fragment size analysis, as shown in the figure below:
What’s more, given that the NuQuant method is incorporated within the library preparation kits, you’re no longer limited in the number of samples you can QC at once. This is an improvement on many other QC processes – with Qubit it’s only possible to run one sample at a time, and Bioanalyzer can only accommodate 11 samples per run. However, NuQuant enables you to QC all samples simultaneously, saving you time and reducing cost.
In addition, the simplicity of the six-minute workflow means that there’s minimal user-user variability; the assay doesn’t need serial dilutions or additional reagents, so there is very little potential for differences in user technique to introduce variability. This leads to more consistent results when compared to other QC methods.
Another key advantage of the NuQuant workflow is that it eliminates sample loss in QC. This is because the output plate can be read directly in a plate reader, so there is no chance of losing valuable library material. This helps you make the most of each sample in your research.
Accurate quantification is also vital in NGS workflows – with better quantification accuracy, it’s possible to normalize library concentration more effectively and therefore generate better-quality data. NuQuant offers a highly accurate quantification method, as shown in the figure below, which highlights the strong correlation (R = 0.97) between the number of reads per sample and NuQuant molar concentration: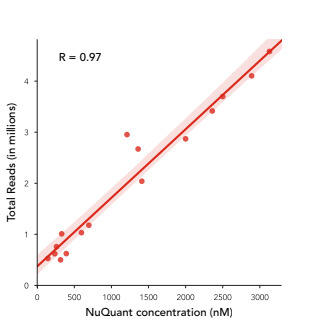
The benefits of optimizing NGS library QC
Optimizing your approach to NGS library QC can make a huge difference to your overall NGS workflow, improving data quality while also saving you time and resources. As such, reliable and efficient QC helps you to accelerate the progress of your research while making your budget go further.
While commonly used QC methods have limitations, innovative new solutions are now available that overcome many of these issues. The NuQuant method is one such example, which has been designed to streamline library preparation, allowing you to accurately QC all libraries simultaneously to reduce the time required. To find out more about NuQuant and review more data supporting this method, download our latest application note today.
About the author
-1.png)
Enrique Neumann
Dr Enrique Neumann is Product and Application Manager, Genomics, at Tecan, Switzerland. He studied Biology at the University of Santiago de Compostela, Spain. During his PhD at the University of Edinburgh, he focused on the molecular processes in plant cells. He joined Tecan in 2015 and focuses on the development and support of genomic applications for Tecan’s liquid handling platforms.